
DockTox: A new tool to support mechanistic toxicology with molecular docking
At ProtoQSAR, we are excited to introduce DockTox, an online tool designed to streamline the molecular docking process for toxicological applications. Developed as part of our contributions to the ONTOX, DockTox represents a key innovation in the move toward animal-free, mechanism-based chemical safety assessment.
Understanding toxicity at the molecular level
DockTox targets a critical concept in predictive toxicology: the Molecular Initiating Event (MIE), the first interaction between a chemical compound and a biological macromolecule that triggers an adverse outcome. Understanding how a compound interacts with these key protein targets is essential for anticipating toxicological effects, especially within the framework of Adverse Outcome Pathways (AOPs).
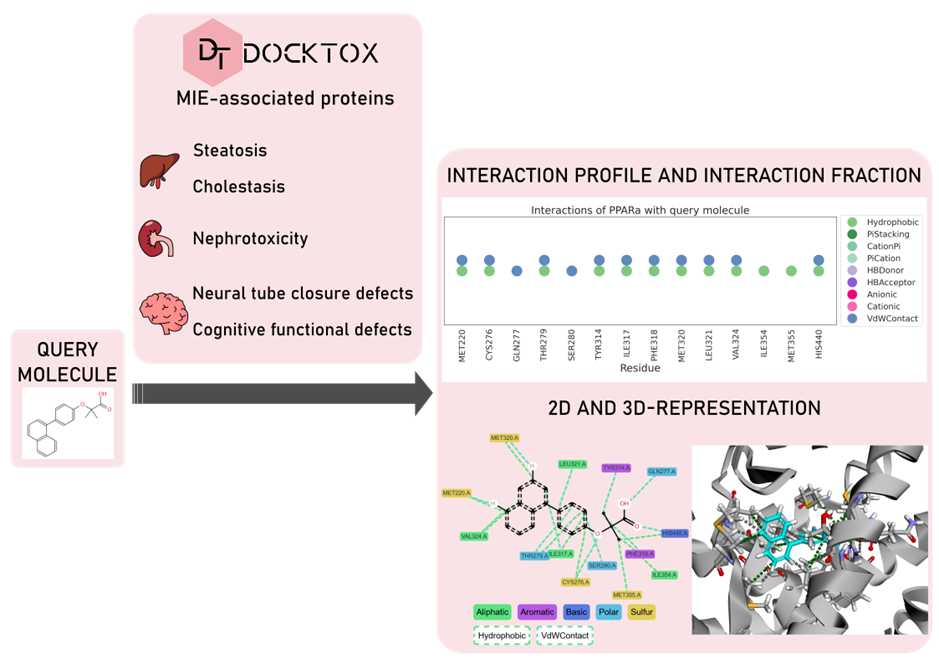
To support this, DockTox offers a fully automated and user-friendly web platform where researchers and toxicologists can screen small molecules against a curated list of proteins associated with MIEs relevant to organ toxicity, including hepatic steatosis, cholestasis, nephrotoxicity, and developmental neurotoxicity.
Why DockTox?
Existing docking platforms often demand prior expertise in protein and ligand preparation, complicating their adoption by users without computational backgrounds. DockTox solves this by automating structure preprocessing, docking setup, and result interpretation. It provides not only binding energy values, but also interaction maps and a unique “interaction fraction” metric (a measure of how well a compound reproduces key binding interactions seen in known active ligands). This metric, combined with binding energy, helps distinguish between likely binders and non-binders.
A generalizable platform
Although the current release of DockTox includes 23 pre-processed proteins relevant to liver, kidney, and brain toxicities, the workflow behind the platform is designed to be easily extendable to other MIE-related proteins. This opens the door for DockTox to support research on broader toxicological endpoints or disease mechanisms involving compound-protein interactions.
Research roots and regulatory relevance
DockTox is one of the core outcomes of ongoing research by our team at ProtoQSAR within the ONTOX consortium and forms part of the doctoral research of our colleague Rita Ortega-Vallbona, whose work focuses on the development of computational tools for the prediction of drug-induced liver injury (DILI), with a particular emphasis on steatosis.
The development of DockTox aligns with the ONTOX vision: building Next Generation Risk Assessment (NGRA) workflows that integrate New Approach Methodologies (NAMs). With its intuitive interface and strong mechanistic foundation, DockTox brings structure-based approaches closer to regulatory acceptance, supporting a tiered weight-of-evidence strategy alongside QSAR, SAR, read-across, and omics-based models.
Get to know DockTox
You can learn more about the development, implementation, and performance evaluation of DockTox in our recently published article:
Ortega-Vallbona, R., et al. (2025). DockTox: Targeting molecular initiating events in organ toxicity through molecular docking. Toxicology, 515, 154155.
📖 https://doi.org/10.1016/j.tox.2025.154155
DockTox is freely available for academic users at:
👉 https://chemopredictionsuite.com/DockTox
We invite toxicologists, pharmacologists, and chemical safety experts to explore this tool and consider it as part of their mechanistic assessment strategy. Whether you’re exploring chemical-induced toxicity or studying molecular mechanisms of adverse outcomes, DockTox provides a gateway to mechanistic understanding through molecular docking.

This work was performed in the context of the ONTOX project (https://ontox-project.eu/), which has received funding from the European Union’s Horizon 2020 Research and Innovation programme under grant agreement No 963845. ONTOX is part of the ASPIS project cluster (https://aspis-cluster.eu/).





